IGV genome browser usage
This blog records two customized visualization tracks when using IGV genome browser.
- Make customized genome file;
- Heatmap with DNA methylation array data.
IGV genome file (this is only applicative for versions older than 2.11.9)
When the organism you are studying is not support by the default IGV genome server or you just want to keep using the exact same genome with your data analysis procedure, you can customized the genome file in IGV.
Procedures:
- Download the genomic files you plan to use, both the sequence (.fasta) file and annotation (.gtf) file.
- Index your .fasta and .gtf files. As for fasta file, you can use either the igvtools embeded in the IGV software or samtools index to do this job; for .gtf file, you’d better first use the sort and then the index igvtools. (igvtools mentioned can be found on desktop version IGV in the following panel)
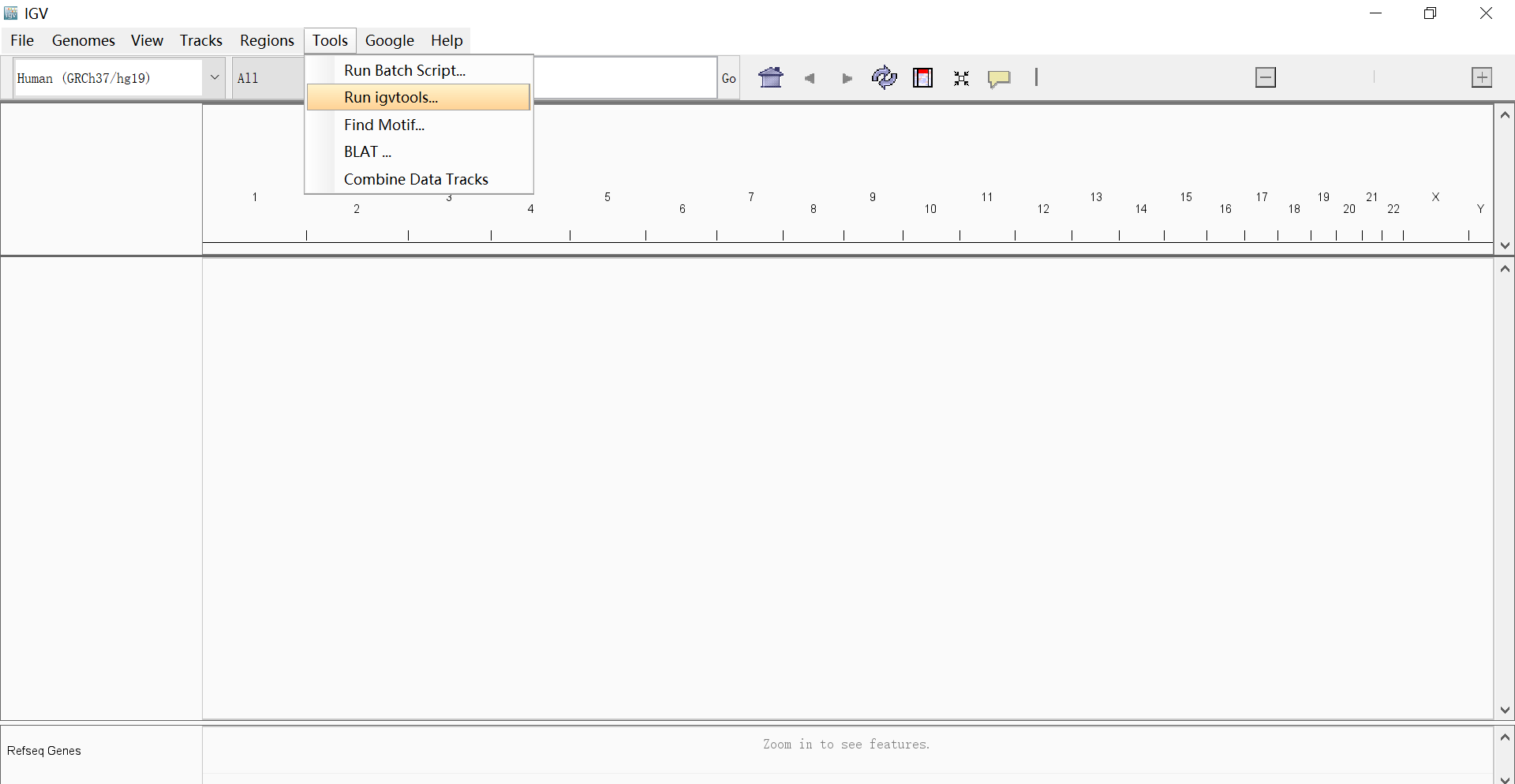
- Create the .genome file
Through theGenomespanel in the main page of IGV to create visualizable genome file.
Genomes->create .genome file
Proper format of DNA methylation data
In order to visualize the DNA methylation data (like beta-value matrix obtained from TCGA) in heatmap format, we need proper files to load in IGV.
Actually, IGV manual provides proper instructions, but I ran into problems in practice, since recording my experience here.
Notes:
The file format need to be like this:
column | description
———|———-
chr | chromosome number (required)
start | start position (required)
end | end position (optional)
probe | description, like a DNA probe name (optional)
data | values to show, like beta-value of DNA methylation probe (required)
And, very importantly, a special index instruction line need to be added in the first line in this file. The content of the line should be like:#columns chr=1 start=2 probe=4 data=5-9
In this line, you need to clarify the column index of the most important features, which is 1-based index.
Warning: pay attention to the blanks between columns, that might be the reason you can’t successfully load your data.
Coding tips for merging two files by a ID column with awk:
1 | awk -F"\t" 'ARGIND==1{a[$1]=$0} ARGIND==2 && ($1 in a) {print $0,"\t",a[$1]}' file1 file2 |