Proteomic sequencing in Alzheimer's disease
This blog is a simple summary for the published proteomic sequencing data in Alzheimer’s disease. The sample number, brain region used for sequencing and specific methods are recorded for further reference.
This summary is based on the researches published untill the blog release date.
Long story short, the samples used for proteome sequencing in AD patients are:
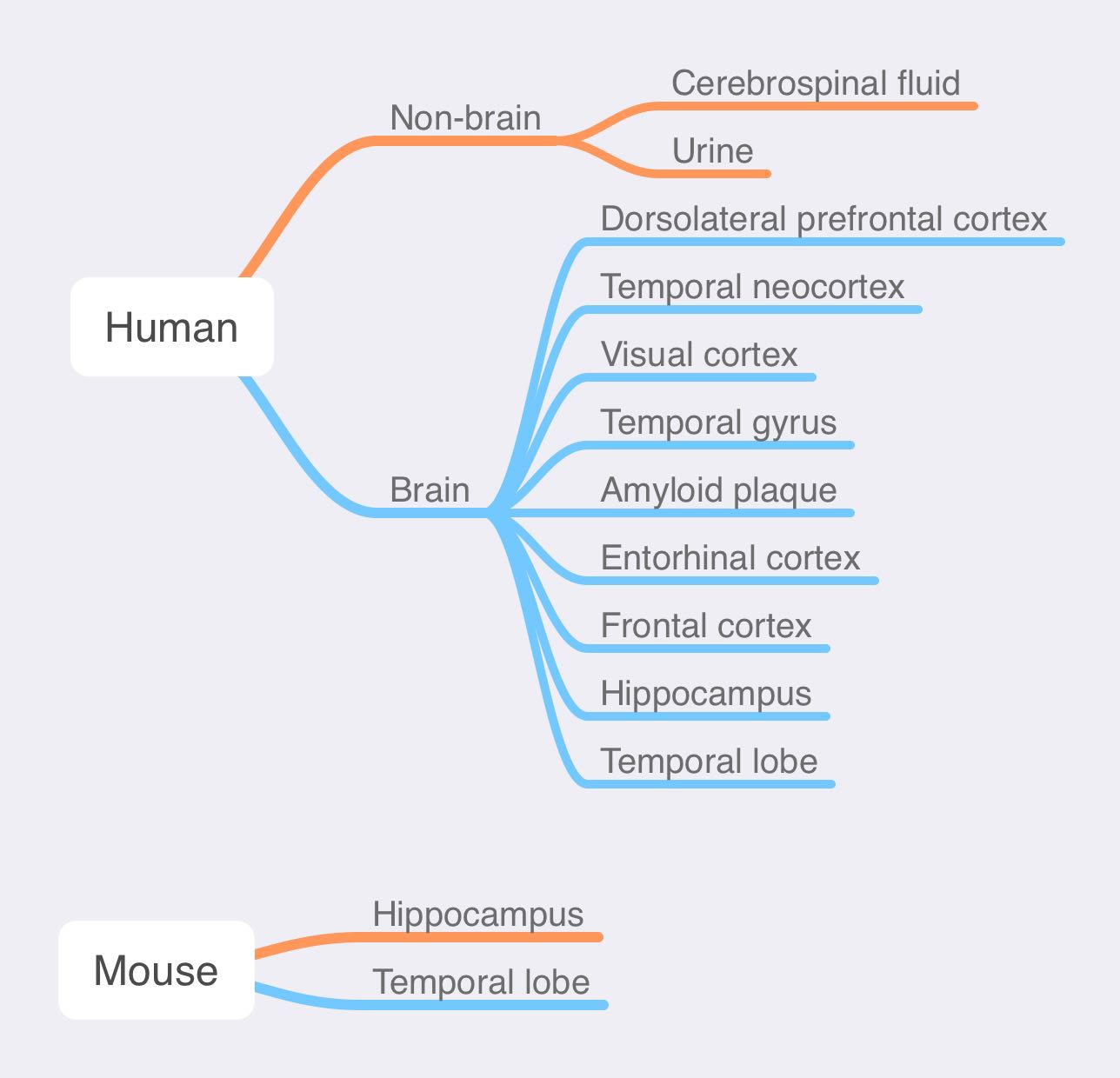
The detailed materials and methods used in each research are as following:
1. Quantification of the Brain Proteome in Alzheimer’s Disease Using Multiplexed Mass Spectrometry. (2014. Journal of Proteome Research.)
- Subjects: Temporal neocortex of Alzheimer’s disease (AD, n=10) patients and non-AD (n=5) Individuals.
- Methods: shotgun mass spectrometry based on a stable isotope dimethyl labeling.
- Results: A total of 827 unique proteins were identified and quantitated. A total of 69 proteins showed different levels (p-value < 0.05) in AD versus control brain samples.
2. Deep proteomic network analysis of Alzheimer’s disease brain reveals alterations in RNA binding proteins and RNA splicing associated with disease. (2018. Molecular Neurodegeneration.)
- Subjects: Dorsolateral prefrontal cortex of symptomatic AD (n = 20), symptomatic AD (AsymAD) (n = 14), and control (n = 13) cases.
- Methods: isobaric tandem mass tag (TMT) mass spectrometry
- Results: 350 proteins had altered levels between AsymAD and AD not caused by changes in specific cell type abundance, potentially reflecting biochemical changes that are associated with cognitive decline in AD. RNA binding proteins were enriched in network modules that correlated with AD pathology.
3. Quantitative proteomics reveals distinct composition of amyloid plaques in Alzheimer’s disease. (2018. Alzheimer’s disease & Dementia.)
- Subjects:
Human: Hippocampal (HC) tissue from three AD donors and three age-matched non-AD donors was used for plaque capture. Hippocampal and temporal lobe (TL) tissue from another 12 donors without neurodegenerative disease was used for human aging brain proteome determination.
Mouse: Whole brains from 6 APP/PS1 model mice and 3 WT mice (all 50 weeks old) were dissected for plaque capture. HC and TL from 4 WT mice in each age group (20, 50, 120 weeks) were dissected for mouse aging brain proteome determination. - Methods: APs and adjacent control regions were collected from fresh-frozen brain sections using laser capture dissection. Tag-labeling coupled high-throughput mass spectra.
- Results: Over 4000 proteins were accurately quantified, and more than 40 were identified as highly enriched in both AD and non-AD APs, including apoE, midkine, VGFR1, and complement C4.
4. Proteomic Atlas of the Human Brain in Alzheimer’s Disease. (2019. Jounral of Proteome Research.)
- Subjects: Nine brain sections: Amygdala (AMY), Caudate Nucleus (CNC), Cerebellum (CBM), Entorhinal Cortex (ECX), Inferior Parietal Lobule (IPL), Middle Frontal Gyrus (MFG), Superior Temporal Gyrus (STG), Thalamus (THA), and Visual Cortex (VCX) in three individuals (The “no tangle” brain (0) contained no observable tangles upon dissection and is not afflicted with Alzheimer’s disease. The “intermediate tangle” brain (+) was categorized as stage III with tangles identified primarily in the entorhinal region. The “severe Tangle” brain (++) falls into the stage VI category with tau aggregates widespread and likely resulting in isocortical destruction.)
- Methods: tandem mass spectrometry
- Results: AD affected tissue exhibited slightly elevated levels of tau protein with similar relative expression to factors associated with the AD pathology.
5. Molecular Network Analysis of the Urinary Proteome of Alzheimer’s Disease Patients. (2019. Dementia & Geriatric Cognitive Disorders.)
- Subjects: Urine samples collected from 18 AD patients and 18 age- and sex-matched cognitively normal controls.
- Methods: Mass spectrometry and semi-quantified with the normalized spectral index method.
- Results: The levels of 109 proteins significantly differed between AD patients and controls. Among these, annotation clusters related to lysosomes, complement activation, and gluconeogenesis were significantly enriched.
6. Comparative profiling of the synaptic proteome from Alzheimer’s disease patients with focus on the APOE genotype. (2019.Acta Neuropathologica Communications.)
- Subjects: AD, APOE3/3, primary visual cortex, n=8; AD, APOE3/4, primary visual cortex, n=11; AD, APOE3/3, superior temporal gyrus, n=7; AD, APOE3/4, superior temporal gyrus, n=9 ; non-demented controls, APOE3/3, primary visual cortex, n=8; non-demented controls, APOE3/4, primary visual cortex, n=5; non-demented controls, APOE3/3, superior temporal gyrus, n=8; non-demented controls, APOE3/4, superior temporal gyrus, n=7.
- Methods: complex 8-plex TMT LC-MS/MS analysis.
- Results: Proteins involved in glutamatergic synaptic signalling and synaptic plasticity are decreased in AD with temporal cortex (which has high levels of pathology) being more severely affected than occipital cortex (which has lower levels of pathology) and APOE4 carriers more affected than APOE3 carriers.
7. Rapamycin regulates cholesterol biosynthesis and cytoplasmic ribosomal proteins in hippocampus and temporal lobe of APP/PS1 mouse. (2019. Journal of Neurological Sciences.)
- Subjects: proteome profles of hippocampus and temporal lobe of wild-type mice, APP/PS1 mice and rapamycintreated APP/PS1 mice.
- Methods: HPLC-coupled high-throughput mass spectrometry approach.
- Results: Proteome analysis identified 100 significantly changed (SC) proteins in hippocampus and 260 in temporal lobe in APP/PS1 mice. Among these, 57 proteins in hippocampus and 167 proteins in temporal lobe were rescued by rapamycin.
8. Large-scale proteomic analysis of Alzheimer’s disease brain and cerebrospinal fluid reveals early changes in energy metabolism associated with microglia and astrocyte activation. (2020. Nature Medicine.)
- Subjects and methods: (1) dorsolateral prefrontal cortex (DLPFC) tissue in 44 cases from the Baltimore Longitudinal Study of Aging (BLSA), 178 cases from the Banner Sun Health Research Institute (Banner), 166 cases from the Mount Sinai School of Medicine Brain Bank (MSSB) and 65 cases from the Adult Changes in Thought Study (ACT), for a total of 453 control, asymptomatic AD (AsymAD) and AD brains; MS-based proteomics using label-free quantitation (LFQ). (2) 340 DLPFC brain tissues from a community-based aging cohort, the Religious Orders Study and Memory and Aging Project (ROS/MAP); with a different MS-based protein quantification approach using isobaric multiplex tandem mass tags (TMTs). (3) control and AD brain tissue from the temporal cortex in a separate set of 111 brains from the Mayo Clinic and control, AsymAD and AD brain tissue from precuneus in the same set of brains from the BLSA; LFQ-MS. (4) DLPFC tissues from Johns Hopkins in 84 individuals aged 30 to 69 years; LFQ-MS. (5) 331 DLPFC tissues from control individuals and those with AD, amyotrophic lateral sclerosis (ALS), frontotemporal lobar degeneration with TDP-43 pathology (FTLD-TDP), progressive supranuclear palsy (PSP), corticobasal degeneration (CBD), Parkinson’s disease and Parkinson’s disease dementia (PD/PDD) and multiple systems atrophy (MSA); LFQ-MS. (6) cerebrospinal fluid (CSF) from two separate cohorts: one cohort of 297 individuals consisting of controls and patients with AD (Cohort 1) and a second cohort of 96 individuals classified into control, AsymAD and AD (Cohort 2); TMT-MS.
- Results: A protein network module linked to sugar metabolism emerged as one of the modules most significantly associated with AD pathology and cognitive impairment. Proteins from this module were elevated in cerebrospinal fluid in early stages of the disease.
9. Integrated proteomics reveals brain-based cerebrospinal fluid biomarkers in asymptomatic and symptomatic Alzheimer’s disease. (2020. Science Advance.)
- Subjects: cerebrospinal fluid (CSF) : discovery cohort: 20 cognitively normal individuals and 20 individuals with AD; CSF replication 1 included samples from 32 healthy controls, 31 individuals with AsymAD, and 33 individuals with AD. CSF replication 2 contained 147 control and 150 AD samples.
- Brain tissue: DLPFC 10 healthy control, 10 PD, 10 AD/PD, and 10 AD cases. The replication brain cohort included 19 cases identical to the discovery cohort (10 control and 9 AD cases), as well as 8 AsymAD cases unique to this cohort.
- Methods: multiplex tandem mass tag (TMT) labeling mass spectrometry.
- Results: Multiplex mass spectrometry identified ~3500 and ~12,000 proteins in AD CSF and brain, respectively. Synaptic and metabolic panels were decreased in AD brain but increased in CSF, while glial-enriched myelination and immunity panels were increased in brain and CSF.
10. Deep Multilayer Brain Proteomics Identifies Molecular Networks in Alzheimer’s Disease Progression. (2020. Neuron.)
- Subjects: Frontal cortical samples of 100 human cases and selected 90 samples of high quality for profiling the whole proteome and phosphoproteome in 5 equal groups: (1) controls with a low pathology of plaques and tangles (LPCs), (2) controls with high Ab pathology but no detectable cognitive defects (HPCs), (3) mild cognitive impairment (MCI) with Ab pathology and a slight but measurable defect in cognition, (4) late-stage AD with high pathology scores of plaques and tangles, and (5) progressive supranuclear palsy (PSP), another neurodegenerative disorder of tauopathy.
- Methods: Multiplexed TMT method and two-dimensional LC/LC-MS/MS.
- Results: Highlighting 173 protein changes in 17 pathways. Determine 15 Ab-correlated proteins (e.g., MDK, NTN1, SMOC1, SLIT2, and HTRA1). Multi-omics integration prioritizes AD-related molecules and pathways, including amyloid cascade, inflammation, complement, WNT signaling, TGF-b and BMP signaling, lipid metabolism, iron homeostasis, and membrane transport.
11. Proteomic and Transcriptomic Analyses Reveal Pathological Changes in the Entorhinal Cortex Region that Correlate Well with Dysregulation of Ion Transport in Patients with Alzheimer’s Disease. (2021. Molecular Neurobiology.)
- Subjects: AD entorhinal cortex (EC) samples and non-AD control EC samples. (RNA-seq: 7 AD, 18 control; proteome: 4 AD, 4 control.)
- Methods: Transcriptome based on the Illumina RNA-seq and proteome based on TMT-labelling.
- Results: 57 genes were detected both in transcriptome and proteome data, including 51 in similar altering trends (7 upregulated, 44 downregulated) and 6 in inverse trends when compared AD vs. control.
12. Quantitative proteomic analysis of the frontal cortex in Alzheimer’s disease. (2020. Neurochemistry.)
- Subjects: frontal cortex from AD patients with corresponding age-matched brain samples.
- Methods: Tandem mass tags (TMT) approach coupled to highresolution mass spectrometry.
- Results: LC-MS/MS analysis carried out on an Orbitrap Fusion Lumos Tribrid mass spectrometer led to identification of 8,066 proteins. Of these, 432 proteins were observed to be significantly altered (>1.5 fold) in their expression in AD brains.
References
- https://pubs.acs.org/doi/10.1021/pr401202d
- https://molecularneurodegeneration.biomedcentral.com/articles/10.1186/s13024-018-0282-4
- https://pubs.acs.org/doi/10.1021/acs.jproteome.9b00004
- https://www.karger.com/Article/FullText/496100
- https://actaneurocomms.biomedcentral.com/articles/10.1186/s40478-019-0847-7
- https://www.jns-journal.com/article/S0022-510X(19)30083-8/fulltext
- https://www.nature.com/articles/s41591-020-0815-6
- https://n.neurology.org/content/94/15_Supplement/4738
- https://alz-journals.onlinelibrary.wiley.com/doi/abs/10.1002/alz.037231
- https://link.springer.com/article/10.1007/s12035-021-02356-3
- https://onlinelibrary.wiley.com/doi/10.1111/jnc.15116